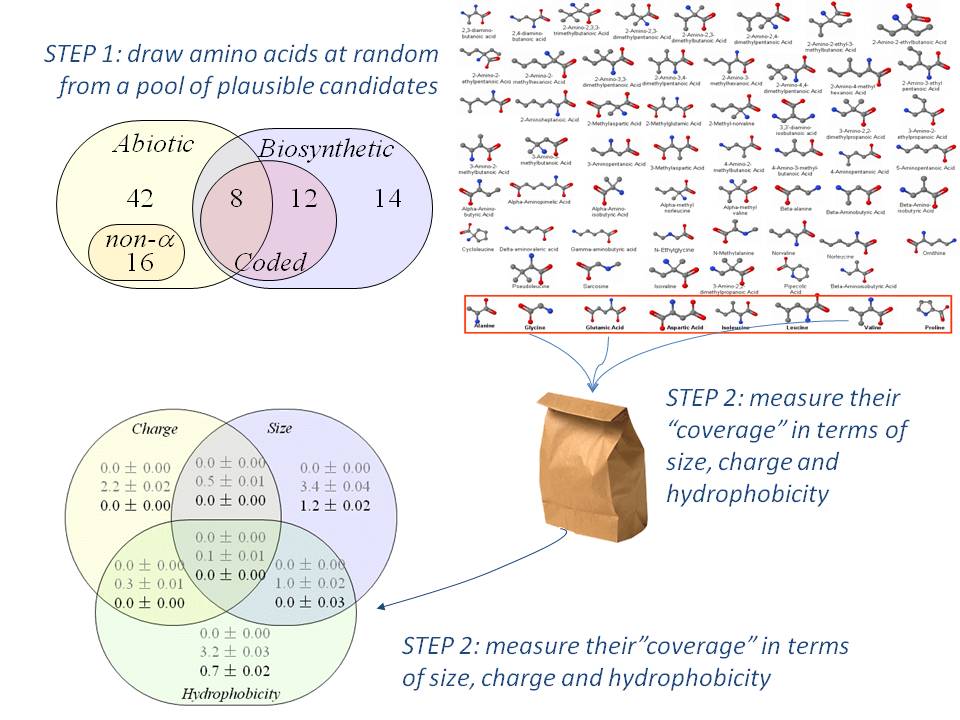
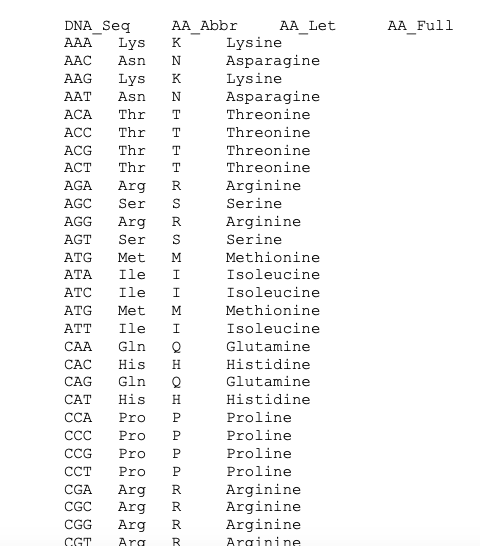
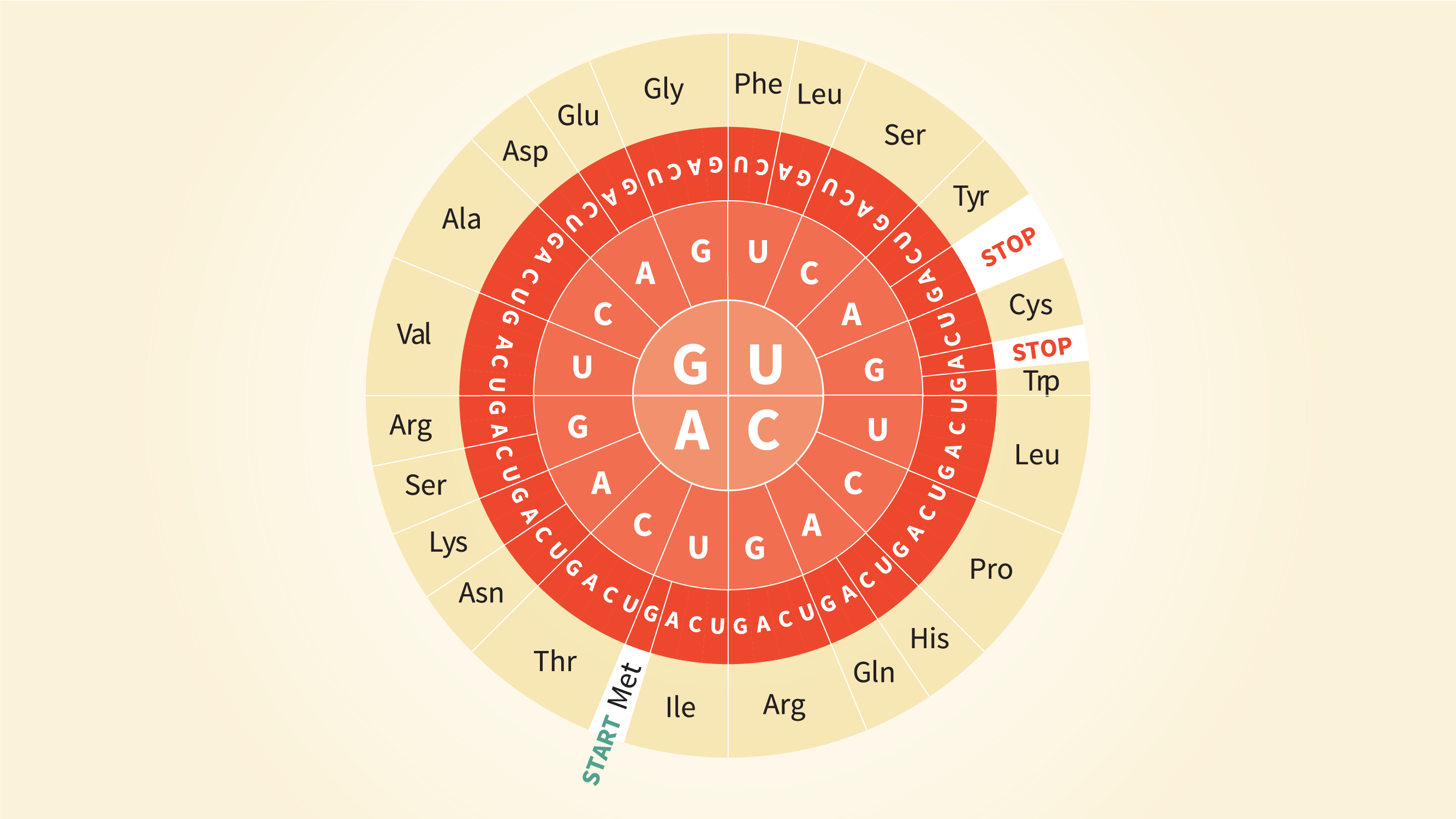
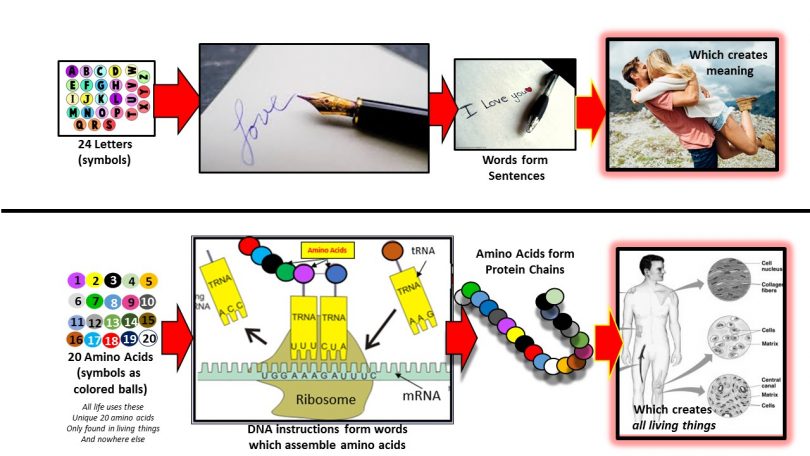
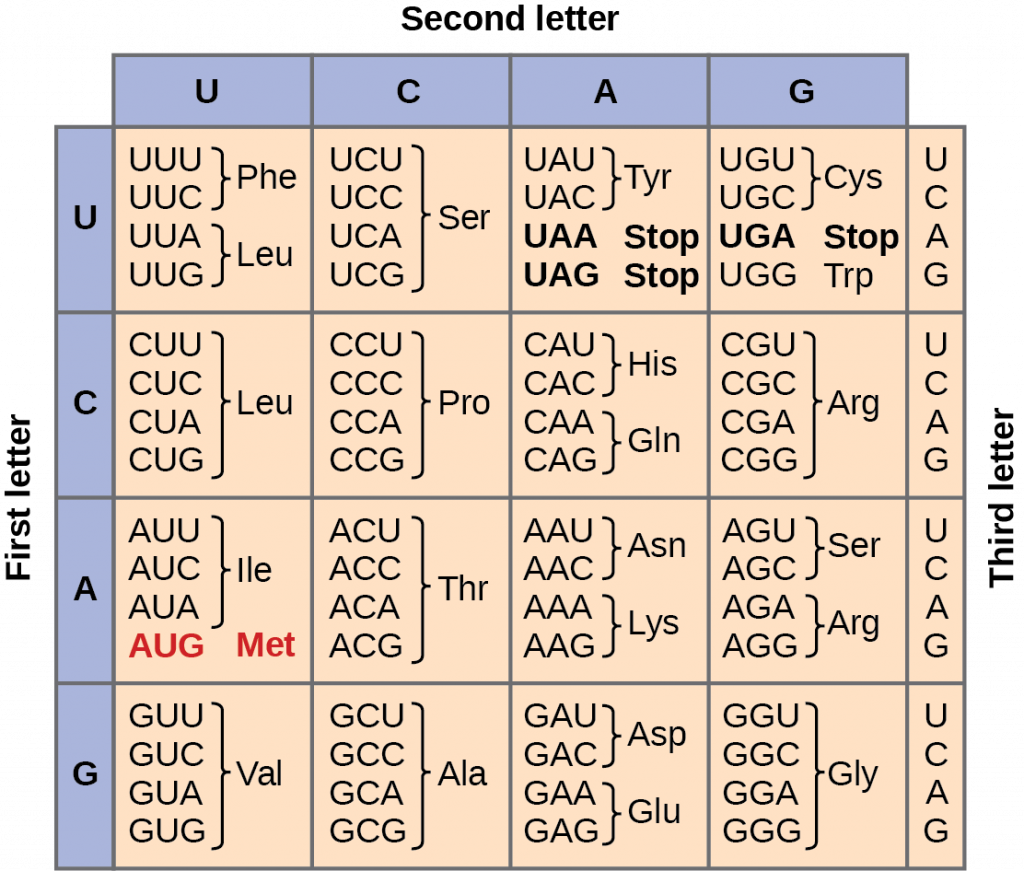
Scientists re-code a whole genome to develop unique species | by Philipp Markolin | Advances in biological science | Medium
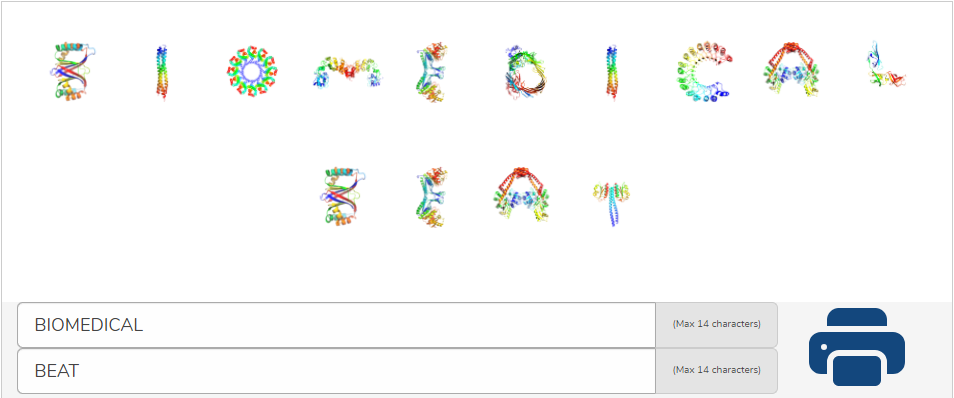
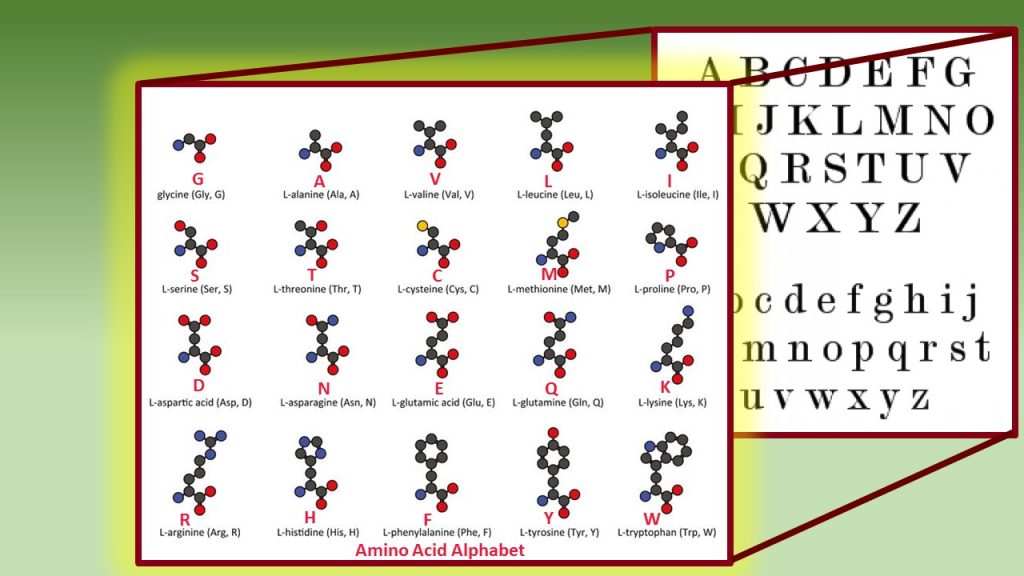
See Your Name With Our Interactive Protein Alphabet! – Biomedical Beat Blog – National Institute of General Medical Sciences

Schemes for reducing amino acid alphabet based on the BLOSUM50 matrix... | Download Scientific Diagram
![PDF] Simplified amino acid alphabets for protein fold recognition and implications for folding. | Semantic Scholar PDF] Simplified amino acid alphabets for protein fold recognition and implications for folding. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/eda832c425dd213a76859b0f06347f1687b647e3/2-Figure1-1.png)
PDF] Simplified amino acid alphabets for protein fold recognition and implications for folding. | Semantic Scholar

Research progress of reduced amino acid alphabets in protein analysis and prediction - ScienceDirect

A reduced amino acid alphabet of two-clusters (AMWLYCFIV-PGHTSDEKNQR)... | Download Scientific Diagram

Genes | Free Full-Text | Improved Large-Scale Homology Search by Two-Step Seed Search Using Multiple Reduced Amino Acid Alphabets
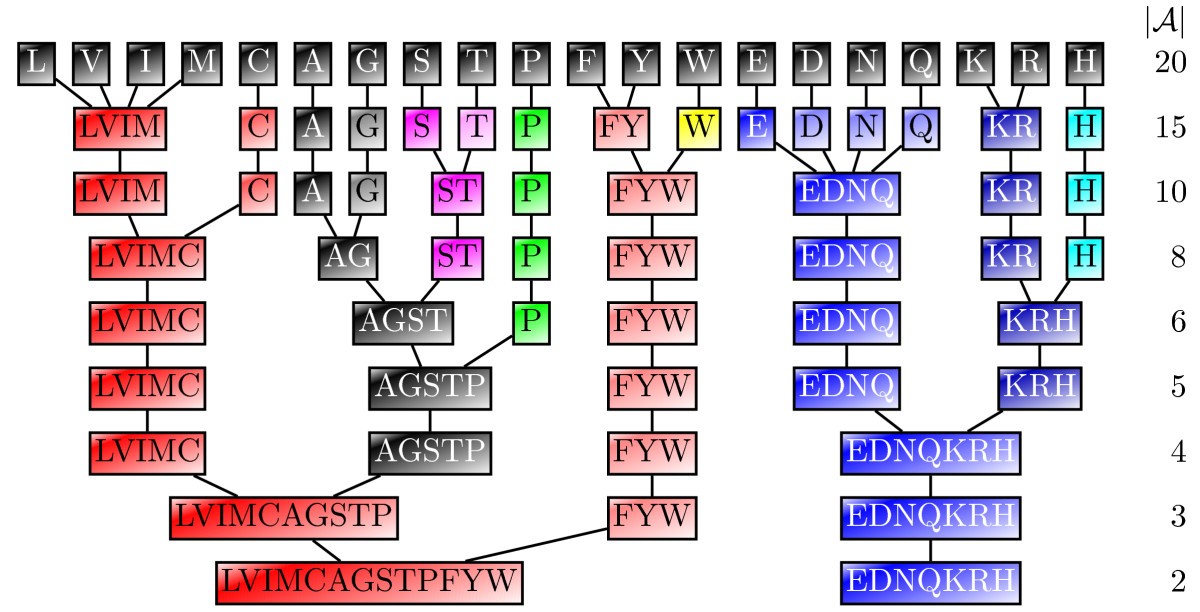
![PDF] On reduced amino acid alphabets for phylogenetic inference. | Semantic Scholar PDF] On reduced amino acid alphabets for phylogenetic inference. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/7bee581280a9a6f843f9e349f2660119e4c9f4af/4-Table1-1.png)